 |
THE PRE-COLUMBIAN PEOPLING OF AMERICA: BY THE Y CHROMOSOME
Abstract
Past human history has been vastly investigated by many distinct disciplines. The first peopling of America has been a focus of an intensive debate among scholars. Genetics has brought a new light to this history but also more discussion and doubts. Even among geneticists, distinct tools have been used and sometimes gave different answers. Now, with the blooming of the genomic era, a large set of genetic data is available and new analyses are being performed with aims to unveil the human past. Variation associated to the human Y chromosome is one of the tools which have added new information about the movement of prehistoric males into America. After almost a decade since the first Y chromosome clue about the American prehistory has been published, we have a better picture of this past event. The Y chromosome record tells that the first migrants have come through Beringia sometime between 18,000 and 15,000 years ago in a single major wave. Those first migrants, the Palaeoamericans, gave rise to the three distinct Native American linguistic groups: Amerindians, Na-Dené and Eskimo-Aleuts. The populations in Asia that are more related to the Native Americans live today in South-Central Siberia close to the Altai Mountains. This genetic history agrees well with other genetic markers like mtDNA and also with several evidences from physical anthropology, archaeology and geology.
Genetics and tracing of prehistorical human migrations
Human history has been always a multidisciplinary study and genetics has been recently added to this endeavor. Similar to other disciplines, genetics looks for ancient traces of the human past to draw a picture of our evolutionary prehistory. It has been used with success to investigate the migration of human prehistorical populations. Before the beginning of modern transoceanic journeys in the XV century, humans had already established themselves on all habitable continents. These migrations are thought to have primarily originated in Africa and occurred during the last 100,000 years. Today, populations that still live in the land they inhabited before the XV century are referred to as aboriginal populations. Thus Basques and Germans are aboriginals of Europe, Bantus and Pygmies are aboriginals of Africa, and Native Americans are the aboriginals of America. Most of these populations have few historical records of their origins and, in any case, these do not go back beyond 5,000 years ago.
To investigate prehistoric events, scientists have dealt mostly with vestiges left from ancient cultures (archaeology), fossil bones (physical anthropology), or language (linguistics), but more recently they have used the genetic variability in the DNA of living individuals, which preserves a record of their past (1). In addition, modern techniques of DNA recovery from ancient bones and tissues can provide information on the variation present in ancient times, a field called molecular archaeology. Indeed, the DNA analysis of a 40,000 year old Neandertal bone has suggested that the Neandertals are not our direct ancestors (2).
Questions have traditionally been framed in terms of populations: where and when did the common ancestor of all populations live? What was the route of migration of a population? When did they arrive in their homeland? Population-based questions of this kind assume that a population is a discrete entity that can be followed through time. If so, an individual’s DNA will tend be more similar to that of other individuals from the same population than to that of individuals from different populations. The level of genetic similarity (and dissimilarity) between populations can be arranged along a scale of time. Therefore, the more different two populations are, the older the common ancestor they share. However, populations are not isolated: there are migrations between them and they may fuse. Consequently, it is not easy to draw conclusions about populations, particularly over long periods of time.
The human Y chromosome as a tool for molecular anthropology
The molecular analysis of DNA has allowed an alternative approach to be developed. The classic DNA markers, or Mendelian loci, are located on the autosomes, the chromosome pairs named 1 to 22, and are always inherited by the offspring from the mother and the father in equal contributions. However two special segments of DNA show uniparental inheritance, i.e., they come exclusively from the father or the mother. These segments are the Y chromosome which is transmitted only from fathers to sons, and the mitochondrial DNA (mtDNA) which is only passed from mothers to children. They provide examples of DNA lineages which can be followed over long periods of time. The histories of different lineages may be not identical because of chance or distinct male/female contributions to a present-day population. Habits such as polygyny (many wives for a man), and female transfer from other populations, would be examples of factors leading to distinct histories of males and females found in the same population. Small regions of autosomes also provide lineages that can be followed through time, however, most of studies dealing with human prehistorical migrations has concentrated on mtDNA and the Y chromosome.
The human Y chromosome traces paternal lineages which can be studied through the analysis of nucleotide variations (and other types) that happen along our past generations. A variation originated by mutation in one ancestral male, will be passed to all his descendants through the paternal lineage, i.e., through the father to son genealogy. Thus, analyzing present-day human Y chromosome variation, a reverse evolutionary approach, we are able to trace paternal lineages through time because of the pattern of accumulation and sharing of mutations (Figure 1). Despite the human Y chromosome can tell only about the history of male lineages which left male descendants, it has been suggested by several studies that it has a strong correlation with geography (3,4). The first Y chromosome variation on single nucleotide that could be easily analyzed has only been described in 1994 (5) but several recent works have made available many new nucleotide variations that could be used for population genetic studies (6,7).
The peopling of America
When Europeans arrived in America at the end of the XV century, they found a group of people who were not mentioned in the Bible. These individuals were regarded at the time as the descendants of one of the lost tribes of Israel. A Spanish Jesuit called José de Acosta was the first to suggest, in 1590, that the people now known as Native Americans were descendants of hunters who came from Asia by land a long time ago. This idea is now generally accepted by scientists. In its modern form, it proposes that the ancestors of Native Americans came through Beringia (Figure 2), a land bridge located where the Bering Strait is now found between North America and Siberia, formed when the sea level was lowered during glacial times between 40,000 and 13,000 years ago (8). A likely pathway for further movement into the Americas was through an ice-free corridor which was open irregularly between 36,000 and 20,000 years ago and then completely open after 12,500 years ago when the glaciation ended (8), however an alternative coastal route could have been also used.
There are currently three major debates concerning the pre-Columbian peopling of the Americas:
1) How many distinct groups of individuals made the journey from Asia to Americas?
2) How long ago?
3) Which populations in Asia share the most recent ancestors with the Native Americans?
A former theory, based originally on linguistic data (9), proposes that ancestors of present-day Native Americans came from Siberia in three separate migrations (at different times) giving rise to distinct groups, namely Amerindians (from South, Central and most of North America; e.g. Yanomami), Na-Dené (from North America; e.g. Navajos) and Eskimo-Aleuts (from the tip of North America). According to this theory, the Amerindians are thought to be the only descendants of the first migrants, also called Paleoamericans.
However, new genetic and anthropological data are challenging the Three Migrations theory. Recent findings of ancient bones (~13,500 years old) in South America suggest a distinct and earlier (<14,000 years old) migration of non-Mongoloid people (10), different in appearance from typical East-Asians and present-day Native Americans, that should have become extinct without leaving descendants today. This hypothesis cannot readily be investigated by geneticists as they usually only study the DNA of living people, but may in the future be studied by molecular archaeologists. However analyses of Y chromosome and mtDNA lineages are being used to tell the paternal and maternal histories of the first peopling of the Americas (11). Differently, they tell that present-day Native Americans are all descendants of this ancient Pleistocenic migration (Figure 2).
The mtDNA analysis reveals that most present-day Native Americans belong to only four distinct lineages, which are also found (but are rare) in parts of Asia. Thus these Native American mtDNAs can be traced back to only four mothers, who probably lived in Asia but came to America in a single major wave. Population sources have been proposed in the region around Mongolia and south Siberia, while suggested times for their entry into the Americas range between 30,000 and 15,000 years ago (12).
The Y-chromosomal analysis suggests that most males, at least 90% in South America and 50 to 70% in North America, are descended from a single male (13,14,15). In contrast to the four mtDNA lineages, this lineage, that we name Q3, is found almost exclusively in Native Americans and the few Siberian Eskimo populations carrying it present a low frequency due to back-migration from America. The Q3 lineage therefore probably arose in the Americas or Beringia, derived from lineage Q, the second most frequent in Native Americans. Other previous studies demonstrate that Y chromosomes belonging to the Q lineage are present in populations from Siberia, namely the Kets and Selkups from the Yenissey River Basin and the Altai from the Altai Mountains (15,16). Interestingly, Y chromosomes related to Native American ones (Q and Q3) belong to the P and R lineages that are very uncommon in Asians such as Chinese and Japanese, but are very frequent in Europe. The implication of these studies is that a Y chromosome common ancestor existed long time ago (<30,000 years) in Eurasia which gave rise to most present-day Y chromosomes of Europeans, Kets and Altai in Siberia and also of Native Americans. It would imply an ancient migration route to the Americas coming from Africa and passing through northern Eurasia, Siberia and Beringia (14).
Recent analysis of the Y chromosome (17) suggest now that all three major groups, namely Amerindians, Na-Dené and Eskimo-Aleuts have not entered America by separate migrations but all descend from a single ancestral Paleoamerican population. Other low-frequency Y chromosomes, like lineages C, R1a1 and K (11) may have come through additional minor migration movements but the findings do not support a separate entry for each of the three Native American groups (17). The Y chromosome, as well as mtDNA (12), analysis indicates that most of the Native Americans today are descended from the people coming in a first and major migration movement during the Pleistocene. Molecular dating methods are not yet accurate enough to define precisely the Pleistocene time entry, but recent studies with the Y chromosome have suggested that the first entry into Americas happened between 18,000 and 15,000 years ago (18).
The picture drawn by the Y chromosome can be conciliated with recent findings of physical anthropology. An earlier migration to America (<15,000 years ago) could have brought Siberian people with mongoloid and pre-mongoloid cranial morphology, what would explain several findings of ancient (10) and recent Native Americans (19) with pre-mongoloid traces. Thus according to the Y and mtDNA, the first migrants, the Paleoamericans, were ancestors from both types. Following the geological records of past climate changes (8) by the time of the first migration (<15,000 years ago), the Alberta corridor in North America would be closed, thus a likely coastal route could have been used. This population which migrated south first would give rise to present-day Amerindians. In the end of the Last Glaciation (~12,000 years ago) the Na-Dené migrated to its present distribution in North America, and Eskimo-Aleuts, adapted to the ice, remained in circumpolar regions (Figure 2).
References
1 Cavalli-Sforza LL, Menozzi P and Piazza A (1994) The history and geography of human genes. Princeton University Press, Princeton, New Jersey, USA
2 Krings M, Stone A, Schmitz RW, Krainitzki H, Stoneking M, Paabo S. (1997) Neandertal DNA sequences and the origin of modern humans. Cell 90: 19-30.
3 Quintana-Murci L, Krausz C, Zerjal T, Sayar SH, Hammer MF, Mehdi SQ, Ayub Q, Qamar R, Mohyuddin A, Radhakrishna U, Jobling MA, Tyler-Smith C, McElreavey K. (2001) Y-chromosome lineages trace diffusion of people and languages in southwestern Asia. Am J Hum Genet. 68: 537-542.
4 Rosser ZH, Zerjal T, Hurles ME, Adojaan M, Alavantic D, Amorim A, Amos W, Armenteros M, Arroyo E, Barbujani G, Beckman G, Beckman L, Bertranpetit J, Bosch E, Bradley DG, Brede G, Cooper G, Corte-Real HB, de Knijff P, Decorte R, Dubrova YE, Evgrafov O, Gilissen A, Glisic S, Golge M, Hill EW, Jeziorowska A, Kalaydjieva L, Kayser M, Kivisild T, Kravchenko SA, Krumina A, Kucinskas V, Lavinha J, Livshits LA, Malaspina P, Maria S, McElreavey K, Meitinger TA, Mikelsaar AV, Mitchell RJ, Nafa K, Nicholson J, Norby S, Pandya A, Parik J, Patsalis PC, Pereira L, Peterlin B, Pielberg G, Prata MJ, Previdere C, Roewer L, Rootsi S, Rubinsztein DC, Saillard J, Santos FR, Stefanescu G, Sykes BC, Tolun A, Villems R, Tyler-Smith C, Jobling MA. (2000) Y-chromosomal diversity in Europe is clinal and influenced primarily by geography, rather than by language. Am J Hum Genet. 67: 1526-1543.
5 Seielstad MT, Herbert JM, Lin AA, Underhill PA, Ibrahim M, Vollrath D and Cavalli-Sforza LL (1994) Construction of human Y-chromosomal haplotypes using a new polymorphic A to G transition. Hum Mol Genet 3: 2159-2161
6 Underhill PA, Shen P, Lin AA, Jin L, Passarino G, Yang WH, Kauffman E, Bonne-Tamir B, Bertranpetit J, Francalacci P, Ibrahim M, Jenkins T, Kidd JR, Mehdi SQ, Seielstad MT, Wells RS, Piazza A, Davis RW, Feldman MW, Cavalli-Sforza LL, Oefner PJ (2000) Y chromosome sequence variation and the history of human populations. Nat Genet 26: 358-361
7 Y chromosome consortium (2002) A nomenclature system for the tree of human Y-chromosomal binary haplogroups. Genome Res 12: 339-348
8 Lambeck K, Esat TM, Potter EK. (2002) Links between climate and sea levels for the past three million years. Nature. 419: 199-206
9 Greenberg JH, Turner CG, Zegura SL (1986) The settlement of the Americas: a comparison of the linguistic, dental and genetic evidence. Curr Anthropol 27: 477-498
10 Neves WA, Powell JF, Ozolins EG (1999) Extracontinental morphological affinities of Lapa Vermelha IV, Hominid 1: A multivariate analysis with progressive numbers of variables. Homo 50: 263–282.
11 Schurr TG, Sherry ST. (2004) Mitochondrial DNA and Y chromosome diversity and the peopling of the Americas: evolutionary and demographic evidence. Am J Hum Biol. 16: 420-439.
12 Bonatto SL, Salzano FM (1997) A single and early migration for the peopling of the Americas supported by mitochondrial DNA sequence data. Proc Natl Acad Sci U S A 94: 1866-1871
13 Pena SD, Santos FR, Bianchi NO, Bravi CM, Carnese FR, Rothhammer F, Gerelsaikhan T, Munkhtuja B, Oyunsuren T. (1995) A major founder Y-chromosome haplotype in Amerindians. Nature Genetics 11: 15-16
14 Santos FR, Hutz, MH, Coimbra CEA, Santos RV, Salzano FM, Pena SDJ. (1995) Further evidence for the existence of major founder Y chromosome haplotype in Amerindians. Braz.J.Genet. 18: 669-672.
15 Santos, FR, Pandya, A, Tyler-Smith, C, Pena, SD, Schanfield, M, Leonard, WR, Osipova, L, Crawford, MH and Mitchell, RJ (1999) The central Siberian origin for native American Y chromosomes. Am J Hum Genet 64: 619-628.
16 Karafet, TM, Zegura, SL, Posukh, O, Osipova, L, Bergen, A, Long, J, Goldman, D, Klitz, W, Harihara, S, de Knijff, P, Wiebe, V, Griffiths, RC, Templeton, AR and Hammer MF (1999) Ancestral Asian source(s) of new world Y-chromosome founder haplotypes. Am J Hum Genet 64: 817-831.
17 Tarazona-Santos E, Santos FR. (2002) The peopling of the Americas: a second major migration? Am J Hum Genet 70: 1377-1380.
18 Bortolini MC, Salzano FM, Thomas MG, Stuart S, Nasanen SP, Bau CH, Hutz MH, Layrisse Z, Petzl-Erler ML, Tsuneto LT, Hill K, Hurtado AM, Castro-de-Guerra D, Torres MM, Groot H, Michalski R, Nymadawa P, Bedoya G, Bradman N, Labuda D, Ruiz-Linares A. (2003) Y-chromosome evidence for differing ancient demographic histories in the Americas. Am J Hum Genet. 73: 524-539.
19 Gonzalez-Jose R, Gonzalez-Martin A, Hernandez M, Pucciarelli HM, Sardi M, Rosales A, Van Der Molen S. (2003) Craniometric evidence for Palaeoamerican survival in Baja California. Nature. 425: 62-65.
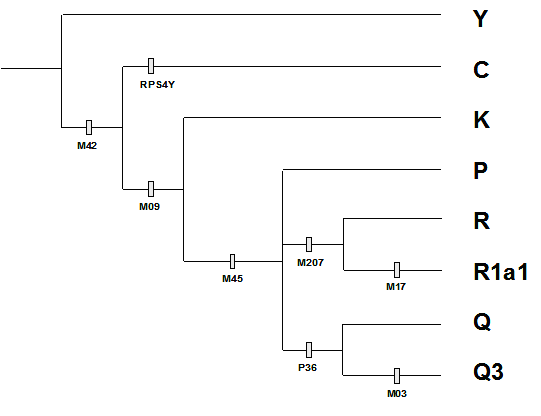
Figure 1 : Tracing Y chromosome lineages (haplogroups) through analyses of accumulation and sharing of variations (gray boxes with loci names).
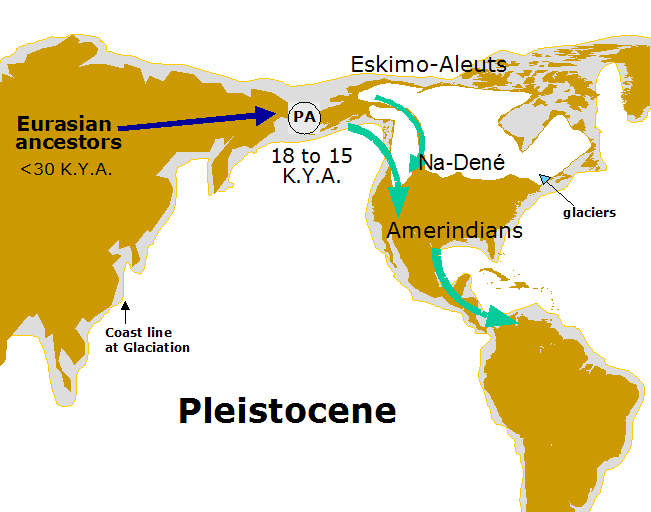
Figure 2 : Pre-Columbian colonization of America by the Y chromosome during the Pleistocene. An ancestral Paleoamerican (PA) population should have crossed Beringia between 18,000 and 15,000 years ago. The Paleoamericans who migrated first to south through the coastline originated the Amerindians. At the end of glaciation (>12,500 years ago) the Na-Dené left the north to south North America and Eskimo-Aleuts remained in the circumpolar area.
[1] Departamento de Biologia Geral, Instituto de Ciências Biológicas, Universidade Federal de Minas Gerais, Belo Horizonte, MG, Brazil. Email: fsantos@icb.ufmg.br